News Journal

News Journal
Cisco Duo wins Best in KLAS again for the second year in a row
Co-author of Allison Norfleet The results of the Best in KLAS Awards 2024: Software and

News Journal
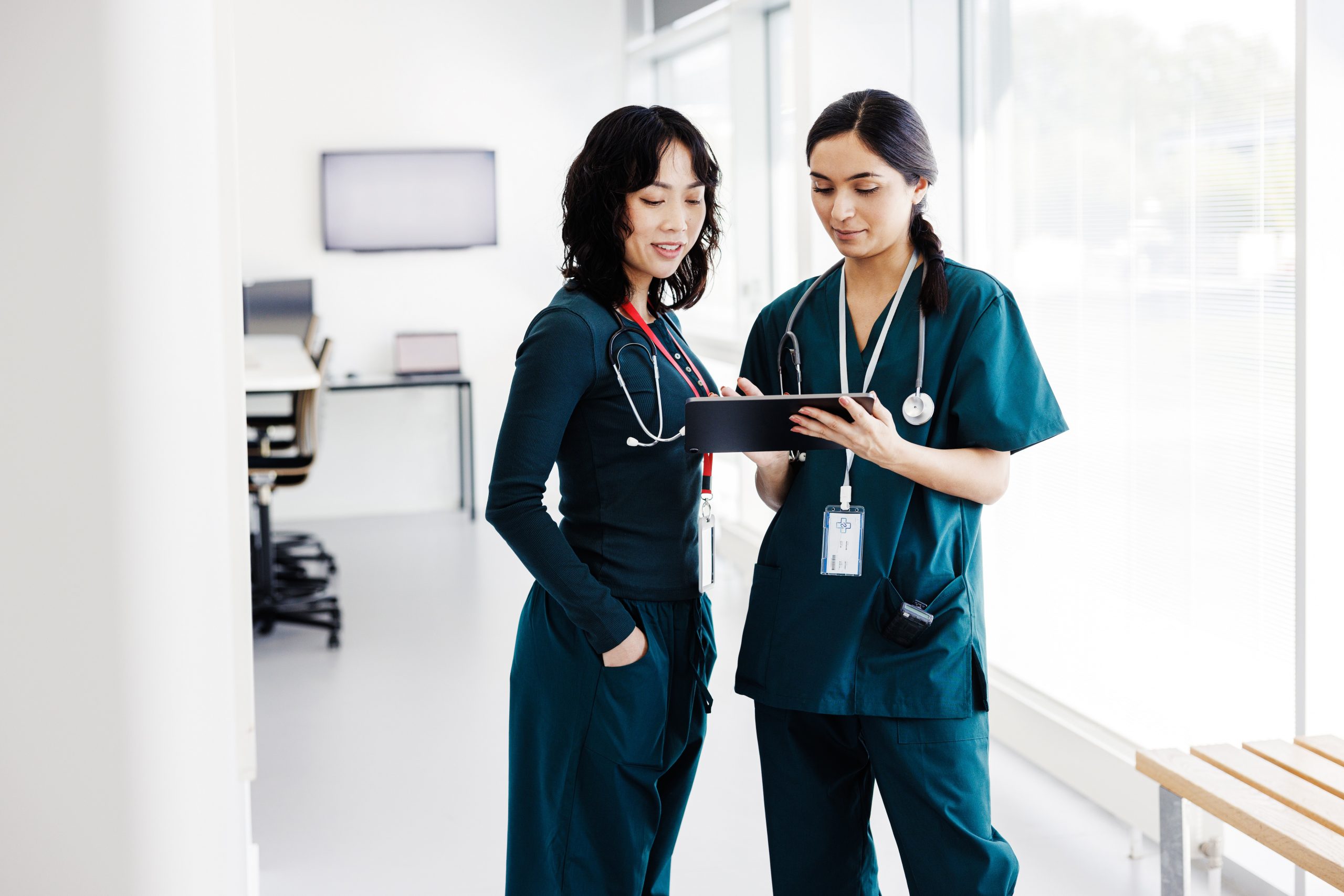
Enhance your Cisco HIMSS24 experience
The Cisco Customer Experience Healthcare Practice helps customers leverage the use of Cisco technology to

News Journal
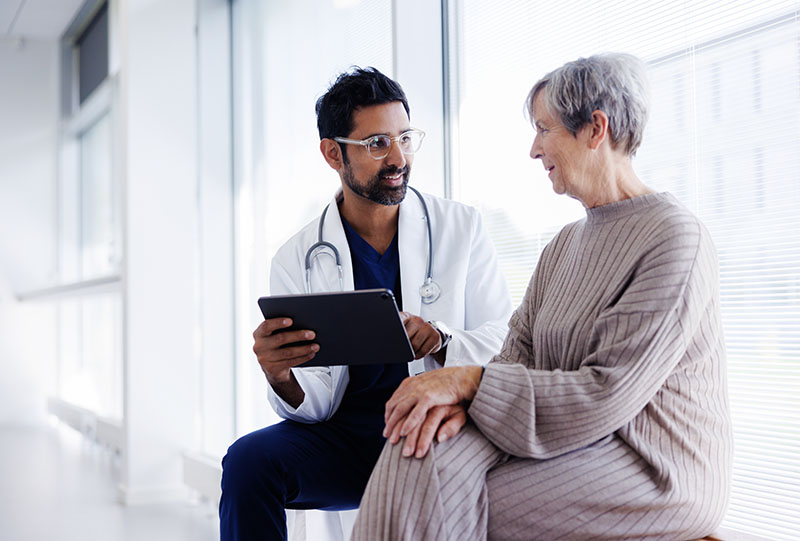
HIMSS 2024 Find out before you go
The future of healthcare is coming into focus! Now more than ever, we believe that

News Journal
How GLP-1 Drug Success Is Transforming Healthcare Revenue: Is Your Organization Ready?
The huge revenue opportunity arising from the recent success of GLP-1 drugs is not just

News Journal
What is the meat and fruit diet?
Many carnivores have begun to add fruit to their diets of animal origin, moving from

News Journal
Primal Essentials giveaway to celebrate 2024
I have a great giveaway for you for the New Year: 6 months of Primal
News From The Web
Co-author of Allison Norfleet
The results of the Best in KLAS Awards 2024: Software and Services the...
Read MoreThe Cisco Customer Experience Healthcare Practice helps customers leverage the use of Cisco technology...
Read MoreThe future of healthcare is coming into focus! Now more than ever, we believe that technology is a critical...
Read MoreThe huge revenue opportunity arising from the recent success of GLP-1 drugs is not just for pharmaceutical...
Read MoreMany carnivores have begun to add fruit to their diets of animal origin, moving from a carnivorous diet...
Read MoreI have a great giveaway for you for the New Year: 6 months of Primal Essentials. These are primal probiotics,...
Read MoreMark’s Apple Primal Essentials Daily Giveaway
Official rules
NO PURCHASE TO ENTER TO WIN. A PURCHASE...
Read MoreResearch of the week
Scythians made of leather out of the skin of his enemies.
5 liters of alkaline water...
Read MoreIn a busy world, efficiency is king. Everyone wants the maximum reward for their efforts in the shortest...
Read MoreThere are certain fundamental inputs that every person needs to be healthy: nutritious food, plenty of...
Read MoreNo posts found